The Personalized NSAID Therapeutics Consortium
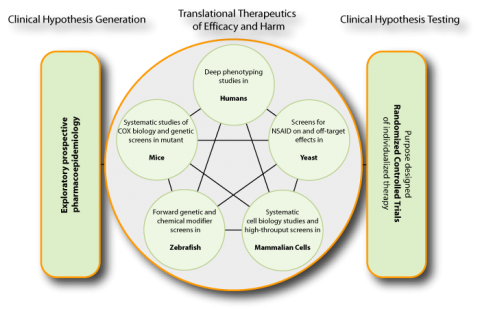
The Personalized NSAID Therapeutics Consortium (PENTACON) is a group of scientists with a wide range of expertise from multiple institutions whose collective aim is to develop a paradigm for the personalization of drug treatment. PENTACON scientists approach the challenge of personalizing chronic drug therapy by focusing on a single class of drugs – nonsteroidal anti-inflammatory drugs (NSAIDs), initially by exploring in detail what factors might contribute to variability in drug response and how that might be reflected by quantitative assays that might predict efficacy or risk. This involves two major challenges: first, the attempt to integrate heterogeneous data – genomics, epigenomics, proteomics, lipidomics, imaging, metabolomics and microbiomics and second, the integration of such data from 5 systems – yeast, mammalian cells, zebrafish, mice and humans.